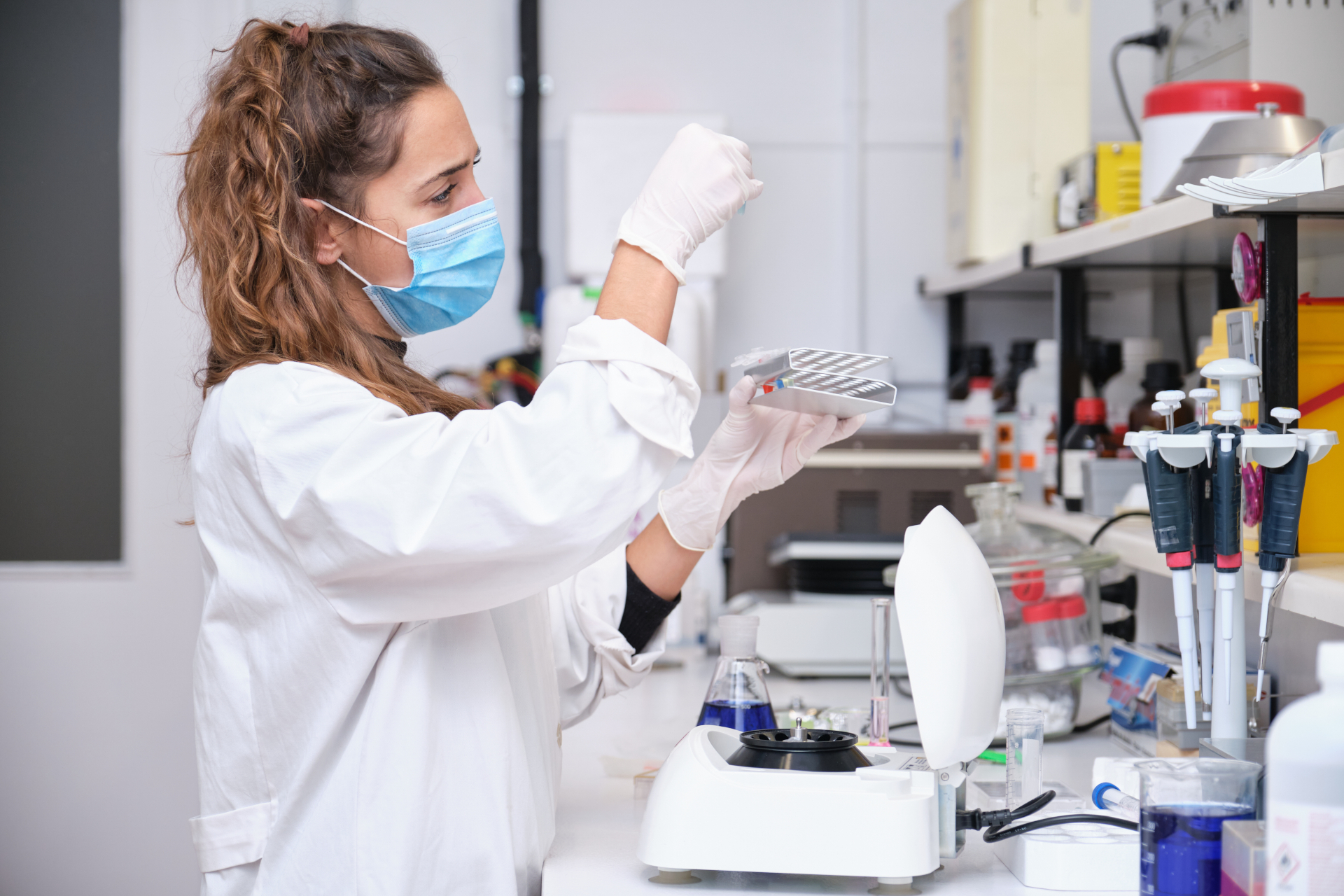
DNA Extraction
EchoLUTION kits for single-step extraction of DNA from various sample types


Stay tuned!
Read the latest news and press releases from BioEcho, download new resources, and see what events we are planning.
Our upcoming events:
EACR 2024
When: 10/6/24, 12:00 PM - 13/6/24, 12:00 PM
Where: Rotterdam, Netherlands | Rotterdam Ahoy, Ahoyweg 10, 3084 BA Rotterdam, Netherlands.
For more information click here.
Webinar: Mastering Nucleic Acid Extraction
When: May 23, 2024 , 4:00 PM
Where: Online.
For more information click here.
Addressing Our Customers' Needs
Animal Breeding & Veterinary Research
Accelerate and simplify your DNA extraction from livestock, poultry, and farm animals.
Academic Research
Get the best out of your nucleic acid extraction for research applications.


Talk to a nucleic acid expert!
Do you have questions on nucleic acid extraction for academic research?